Research Highlights: Hangzhou Institute of Medicine & Affiliated Hospital (December 2025 – February 2026)
Hangzhou Institute of Medicine, Chinese Academy of Sciences
Science
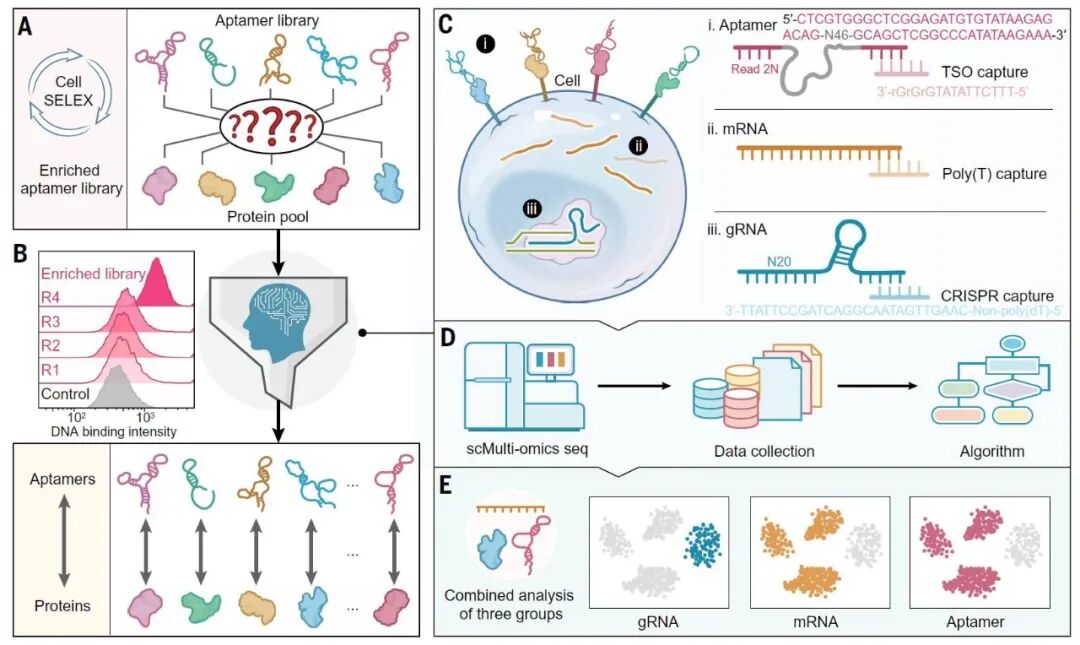
“SPARK-seq: A high-throughput platform for aptamer discovery and kinetic profiling”
Published in Science on 1 January 2026, the study led by Academician Weihong TAN and Prof. Qin WU introduces SPARK-seq, a high-throughput platform that enables simultaneous discovery of cell membrane targets and their specific aptamer probes. Unlike traditional SELEX methods limited by low throughput and poor target identification, SPARK-seq achieves digital precision in membrane protein biomarker discovery, marking a paradigm shift from empirical screening to high-throughput quantitative profiling.
Nature Cell Biology (21 January 2026)
“DNA nanodevices detect an acidic nanolayer on the lysosomal surface”
Published in Nature Cell Biology on 21 January 2026, the collaborative study led by Academician Weihong TAN, Prof. Liping QIU, and Prof. Haoxing XU developed a DNA nanoprobe that anchors to the lysosomal membrane to precisely measure local pH on the lysosomal surface. This work reveals that lysosomes maintain an acidic nanolayer on their membrane surface, providing new insights into how luminal protons may cross membranes to act as signaling molecules in cellular physiology.

Molecular Cancer (22 December 2025)
“The METTL3-YTHDC1 axis mediates architectural RNA m6A modification to modulate the integrity of chromatin TADs in MLLr+AML genome”
Published in Molecular Cancer on 22 December 2025, the study led by Prof. Huacheng LUO, Prof. Wenjuan YU, and Prof. Hanmei LOU reveals that the METTL3-YTHDC1 axis regulates chromatin architectural RNA (arcRNA) m⁶A modification to maintain topologically associating domain (TAD) integrity in MLL-rearranged acute myeloid leukemia. This discovery connects RNA modification to three-dimensional genome organization, offering a new therapeutic target for this high-risk leukemia subtype.

Nature Communications (10 December 2025)
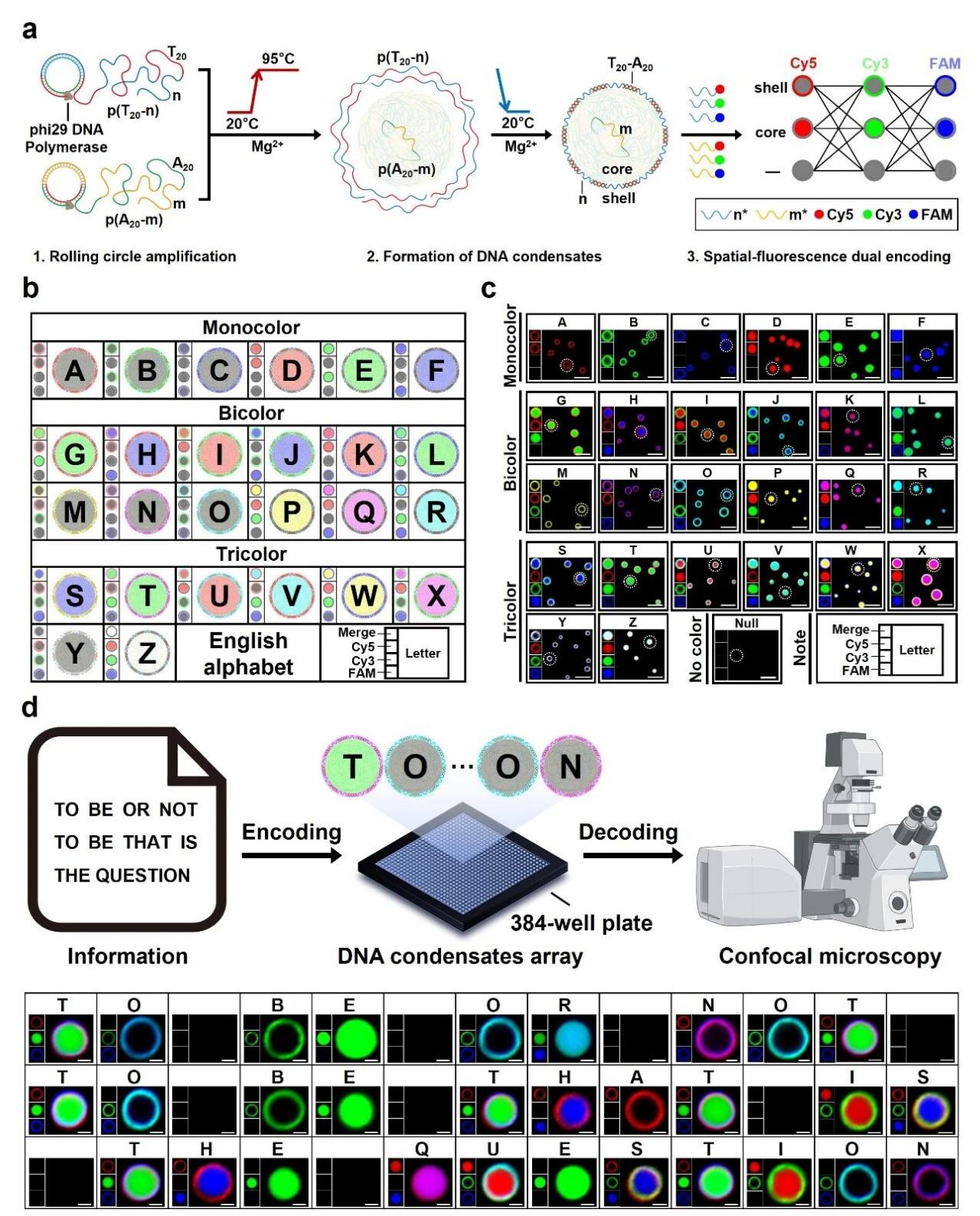
“Hierarchical core-shell DNA condensates enable programmable information storage and encryption”
Published in Nature Communications on 10 December 2025, the study led by Prof. Da HAN and Prof. Lei HE developed a hierarchical core-shell DNA condensate platform that overcomes the static nature of traditional DNA storage. The system enables full information operations, encoding, erasing, rewriting, copying, and restoration, and integrates programmable molecular encryption circuits including logic-gated encryption, biosignal-responsive decryption, time-sensitive encryption, and privilege-based access control.
Nature Communications (11 December 2025)
“Transcytotic transportation of size-controlled nanocarriers into dystrophic skeletal muscle leads to therapeutic outcome in mice”
Published in Nature Communications on 11 December 2025, the study led by Prof. Xiangsheng LIU developed size-controlled nanocarriers that activate transcytosis, a vesicle-mediated transport pathway, in skeletal muscle endothelial cells. The optimized nanoparticles significantly enhanced drug accumulation in dystrophic muscle, improved muscle morphology, and restored motor function in Duchenne muscular dystrophy mouse models, addressing a key delivery challenge in neuromuscular disease therapy.
Nature Communications (24 December 2025)
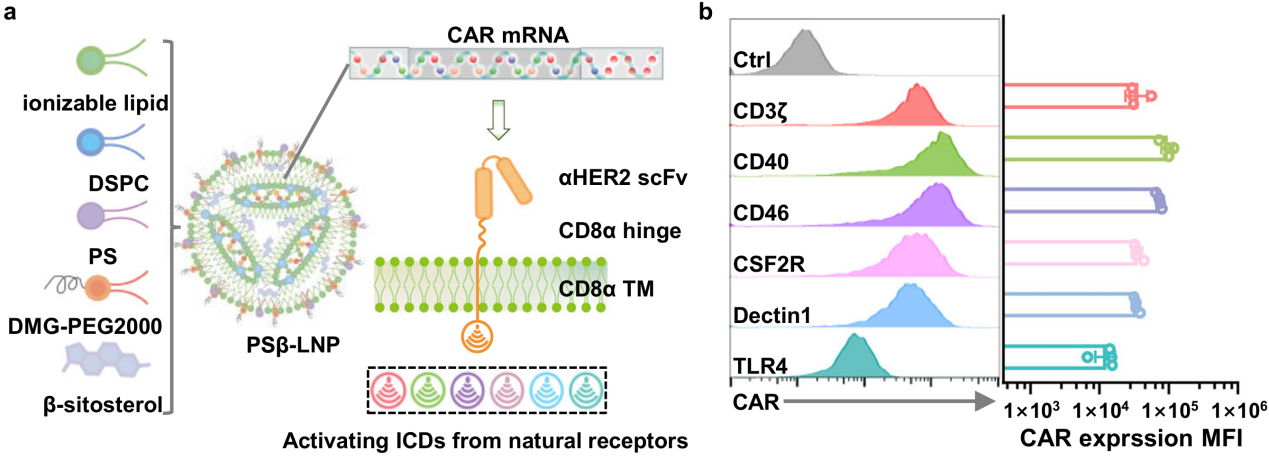
“Intraperitoneal programming of tailored CAR macrophages via mRNA lipid nanoparticle to boost cancer immunotherapy”
Published in Nature Communications on 24 December 2025, the study led by Academician Weihong TAN, Prof. Sitao XIE, and Prof. Xiangsheng LIU screened 36 CAR constructs with different macrophage intracellular domains and achieved in situ intraperitoneal programming of CAR-macrophages using mRNA-LNP systems. This approach bypasses the complex ex vivo manufacturing and viral vector safety concerns of conventional CAR cell therapies, showing robust adaptive immune activation and synergy with PD-1/PD-L1 blockade in solid tumor models.
Nature Communications (7 February 2026)
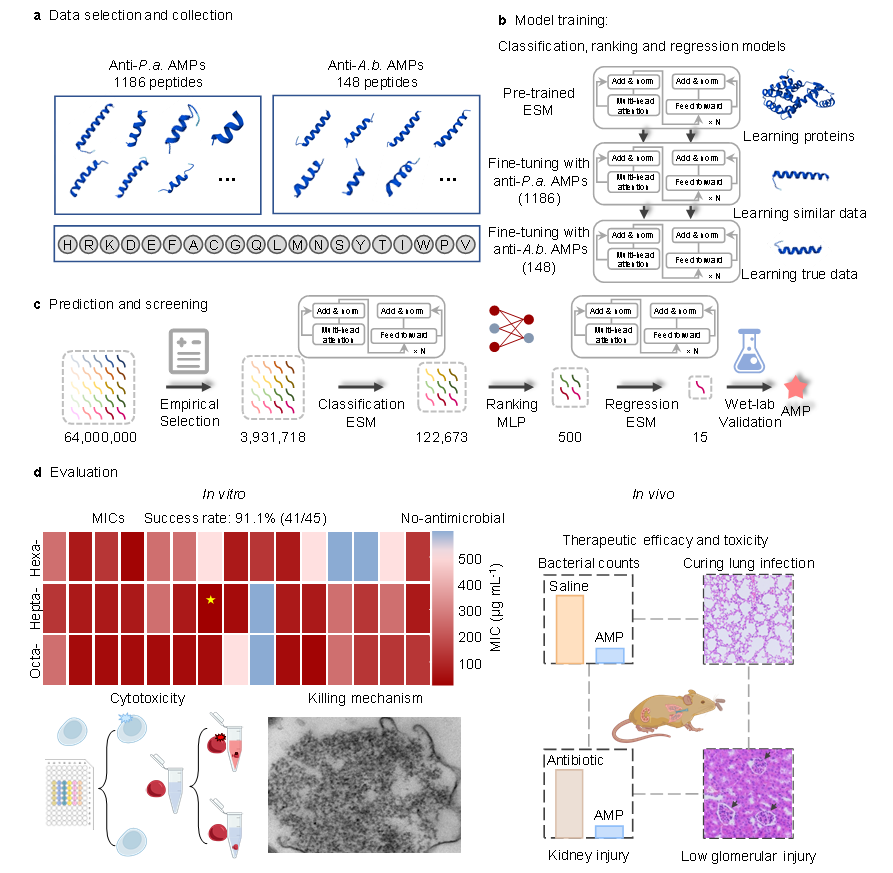
“Discovery of antimicrobial peptides targeting Acinetobacter baumannii via a pre-trained and fine-tuned few-shot learning-based pipeline”
Published in Nature Communications on 7 February 2026, the collaborative study led by Prof. Junjie HUANG, Prof. Jian JI, Prof. Peng ZHANG, and Prof. Junbo ZHAO developed FSLSMEP, a few-shot learning workflow that integrates pre-training with multi-step fine-tuning. The pipeline combines classification, ranking, and regression models to discover antimicrobial peptides against multidrug-resistant Acinetobacter baumannii from peptide libraries, addressing the scarcity of training data for this challenging pathogen.
Journal of the American Chemical Society (December 2025)
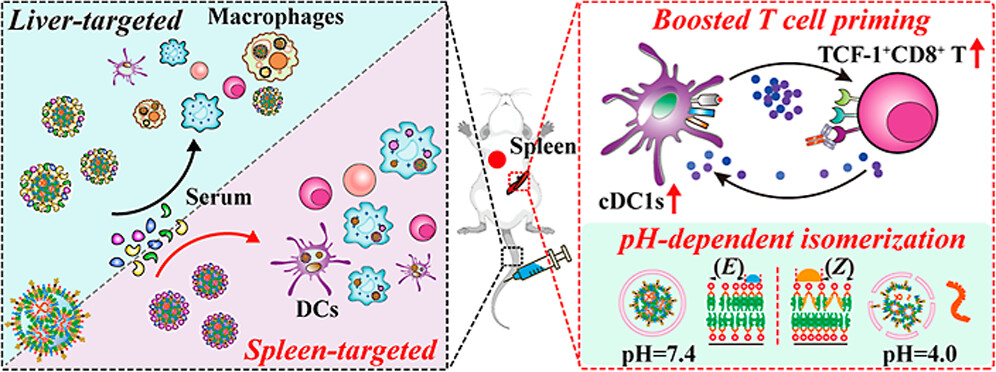
“pH-Isomerizable Acylhydrazone-Based Ionizable Lipids for Spleen-Targeted mRNA Vaccines”
Published in JACS in December 2025, the study led by Prof. Penghui ZHANG developed a new class of ionizable lipids based on acylhydrazone chemistry that undergo pH-dependent isomerization. These AHzILs enable selective mRNA delivery to the spleen, a key organ rich in antigen-presenting cells, overcoming the liver accumulation issue of conventional LNP systems, providing a promising platform for mRNA-based cancer vaccines and immunotherapies.
Angewandte Chemie International Edition (16 January 2026)
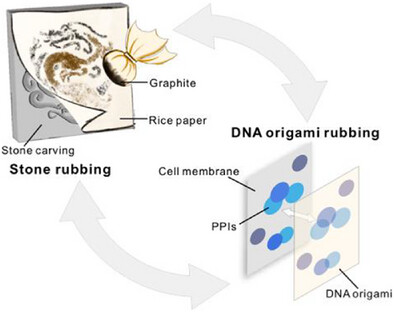
“Spatial Mapping of Membrane Protein Interactions Using a DNA Origami Rubbing”
Published in Angewandte Chemie on 16 January 2026, the study led by Prof. Da HAN, Prof. Pei GUO, and Prof. Qian TANG developed a DNA origami-based “rubbing” strategy that creates spatial imprints of membrane proteins. The method uses DNA origami tiles with precisely spaced recognition sites to map protein nanoscale distribution and capture dynamic changes in response to ligand stimulation. Compatible with antibody-based detection and NGS readout, this platform enables high-throughput mapping of membrane protein interactions in their native environment.
CCS Chemistry (7 January 2026)
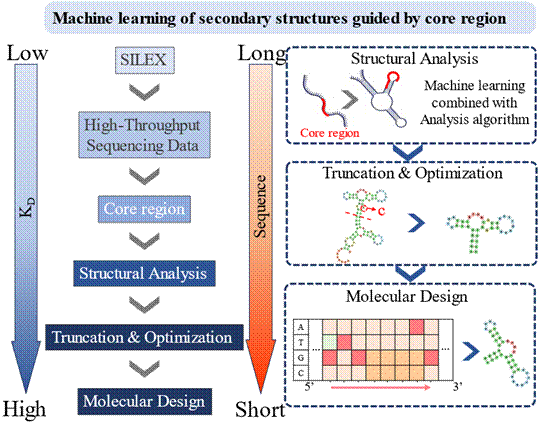
“Single-Round Aptamer Discovery Empowered by Machine Learning: Revealing Structure–Function Principles of Target Binding”
Published in CCS Chemistry on 7 January 2026, the study led by Academician Weihong TAN, Prof. Xiaohong FANG, and Prof. Tao BING developed a machine learning method that directly resolves aptamer secondary structures from single-round screening data without iterative enrichment. This approach enables rational truncation, optimization, and even de novo design of high-affinity aptamers, significantly accelerating the discovery process and revealing structure-function relationships in nucleic acid therapeutics.
Advanced Science (December 2025)
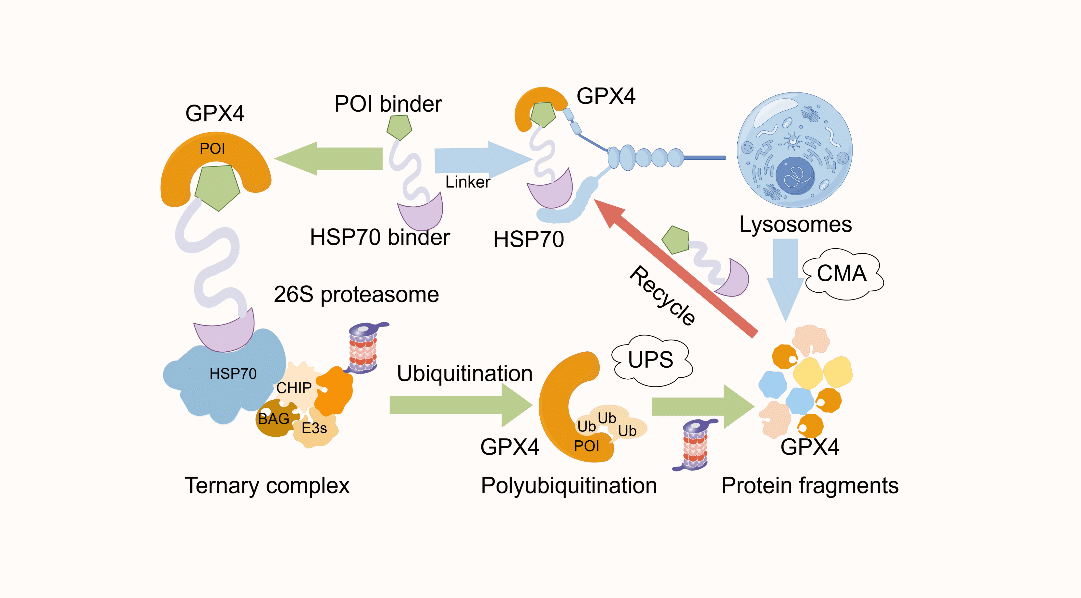
“HSP70 Interactome-Mediated Proteolysis Targeting Chimera (HSP70-PROTAC) for Ferroptosis-Driven Cancer Treatment”
Published in Advanced Science in December 2025, the collaborative study led by Prof. Jiangjiang QIN, Prof. Xuelei MA, and Prof. Fangfang TAO developed HSP70-PROTAC, a novel protein degradation strategy that hijacks the tumor cell-enriched Hsc70 chaperone complex. Unlike conventional PROTACs that rely on CRBN or VHL E3 ligases, this approach maintains degradation activity in CRBN/VHL-deficient cells and offers a new mechanism for ferroptosis-driven cancer therapy.
Affiliated Hospital (Zhejiang Cancer Hospital)
Science Bulletin (February 2026)
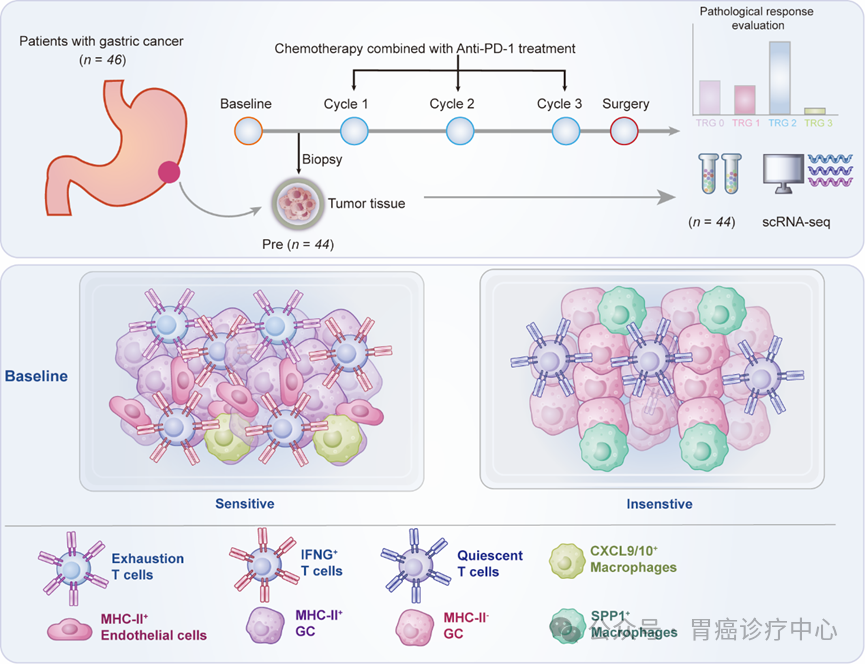
“Comprehensive single-cell profiling uncovers TsMHC-II expression predicting response to neoadjuvant immunotherapy in gastric cancer”
Published in Science Bulletin in February 2026, the study led by Prof. Xiangdong CHENG and Prof. Fan BAI identified tumor cell-derived MHC-II (tsMHC-II) as a predictive biomarker for neoadjuvant immunotherapy response in gastric cancer. In a cohort of 376 patients, tsMHC-II-positive patients achieved a pathological complete response rate of 24.24% with immunochemotherapy versus 6.38% in negative patients. Notably, tsMHC-II specifically predicted immunotherapy benefit (not chemotherapy alone) and outperformed conventional PD-L1 CPS scoring. A prospective validation trial in 30 tsMHC-II-positive patients achieved a 36.67% pCR rate, confirming its clinical utility for patient selection.
Journal of Nanobiotechnology (February 2026)
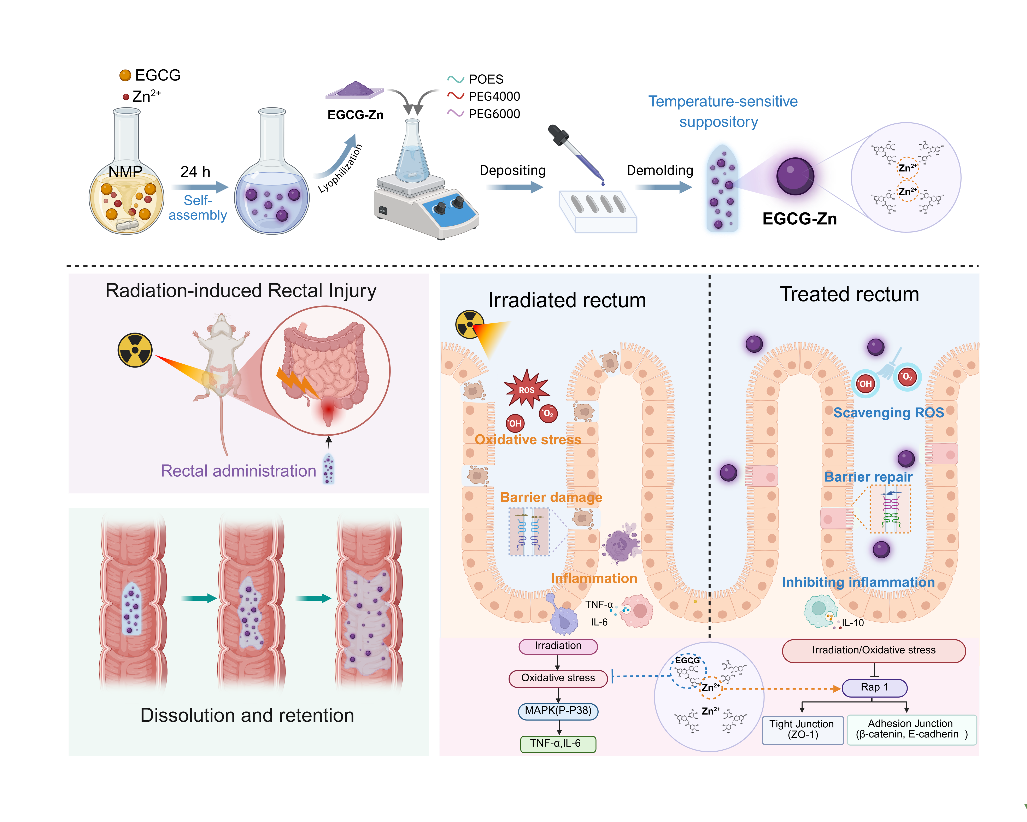
“Rectal delivered nanocomposite suppository against radiation-induced proctitis”
Published in Journal of Nanobiotechnology in February 2026, the study led by Prof. Ji ZHU developed an EGCG-Zn metal-polyphenol nanocomposite formulated as a thermosensitive rectal suppository. The system stabilizes EGCG through Zn²⁺ coordination, enabling sustained local release and retention in rectal mucosa. Mechanistically, it suppresses MAPK-mediated inflammation, reduces oxidative stress, and activates Rap1 signaling to restore tight junction proteins and barrier integrity. This approach breaks the “oxidative stress-inflammation-barrier disruption” cycle, offering a convenient, patient-friendly intervention for radiation-induced proctitis.
Lancet Oncology (February 2026)
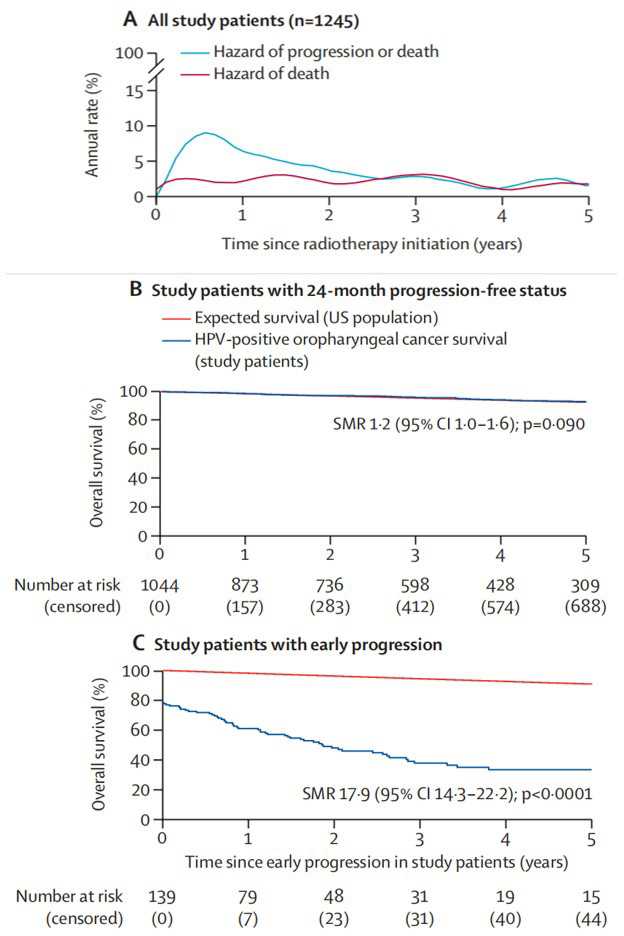
“Evaluation of progression-free status at 24 months as a surrogate endpoint for overall survival in patients with human papillomavirus-positive oropharyngeal cancer: a retrospective cohort study”
Published in The Lancet Oncology in February 2026, this collaborative study led by Dr. Caineng CAO and Prof. Nancy Y. LEE analyzed 1,245 patients with HPV-positive oropharyngeal cancer treated with modern radiotherapy. The study demonstrated that patients remaining progression-free at 24 months (PFS24) achieve subsequent survival rates comparable to the general population. This establishes PFS24 as a potential surrogate endpoint that could reduce trial duration from over a decade to 2-3 years, accelerating the development of new therapies for this favorable-prognosis patient population.
Drug Resistance Updates (December 2025)
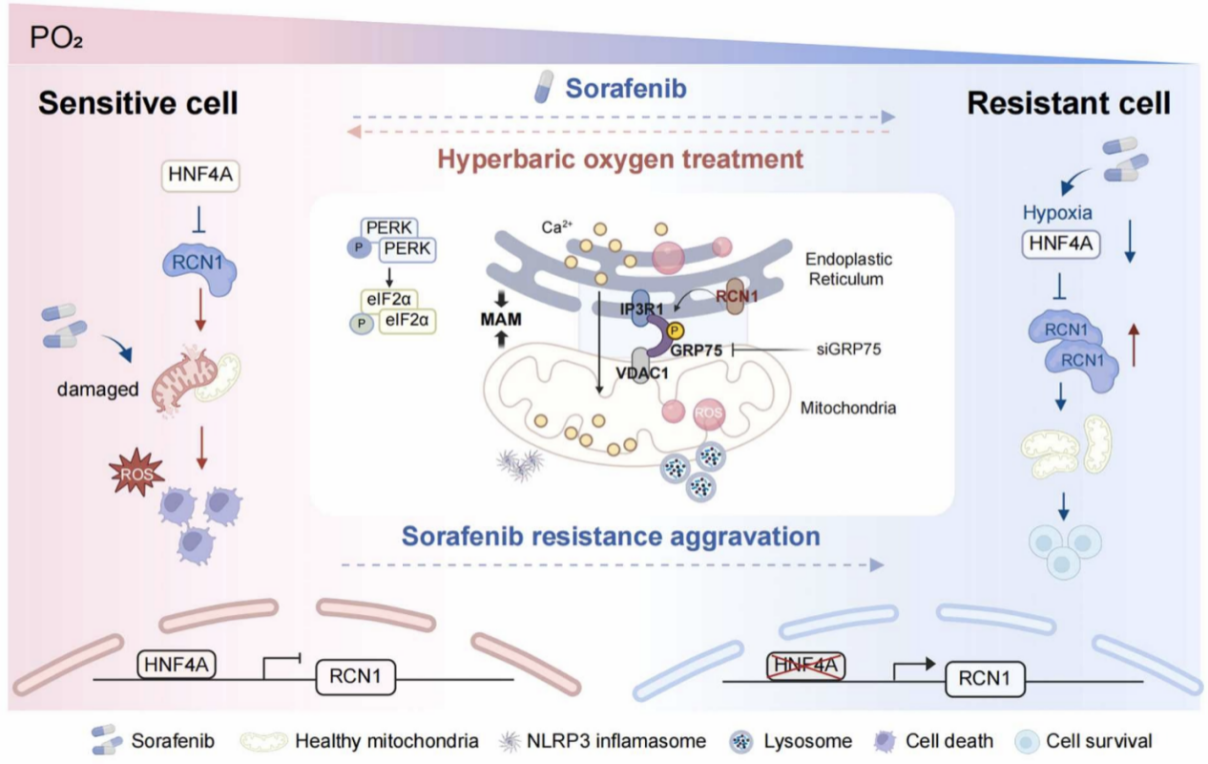
“Hyperbaric oxygen targets RCN1 to modulate ER-mitochondria crosstalk and ameliorate sorafenib resistance in hepatocellular carcinoma”
Published in Drug Resistance Updates in December 2025, the study led by Prof. Weifeng HONG revealed that hyperbaric oxygen therapy reverses sorafenib resistance in hepatocellular carcinoma. Mechanistically, tumor hypoxia suppresses HNF4A and upregulates RCN1, disrupting IP3R1-GRP75-VDAC1 complex assembly and impairing ER-mitochondria calcium signaling. Hyperbaric oxygen restores HNF4A expression, downregulates RCN1, and re-establishes ER-mitochondria contacts, triggering ER stress and mitochondrial dysfunction. This “HBO-HNF4A-RCN1-ER/mitochondria calcium” axis provides a new combination strategy to overcome tyrosine kinase inhibitor resistance in liver cancer.
Advanced Science (February 2026)
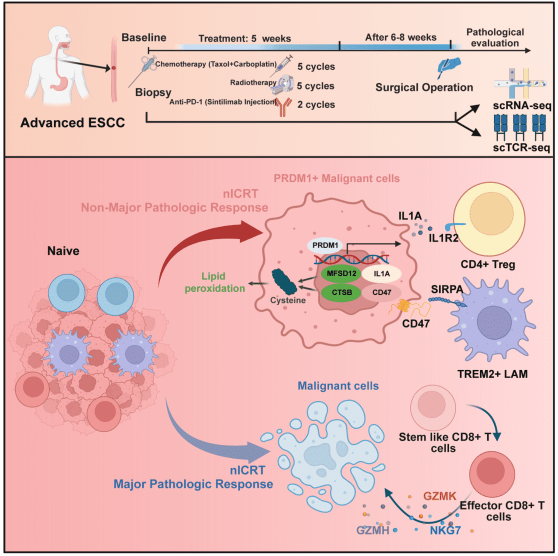
“PRDM1+ Malignant Cells Mediate an Immunosuppressive Landscape and Resistance to Neoadjuvant Chemoradiotherapy and Immunotherapy in Esophageal Squamous Cell Carcinoma”
Published in Advanced Science in February 2026, the study led by Prof. Dijian SHEN identified PRDM1-positive malignant epithelial cells as key drivers of treatment resistance in esophageal squamous cell carcinoma. These cells exhibit lipid peroxidation features mediated by PRDM1 activation of CTSB and MFSD12, making them ferroptosis-sensitive. Importantly, they recruit immunosuppressive regulatory T cells via IL1A-IL1R2 signaling and activate TREM2+ macrophages through CD47-SIRPA, creating an immunosuppressive microenvironment. This work identifies PRDM1 as a resistance-associated factor and suggests targeting ferroptosis or PRDM1-mediated immunosuppression as strategies to overcome treatment resistance.
Journal for ImmunoTherapy of Cancer (5 December 2025)
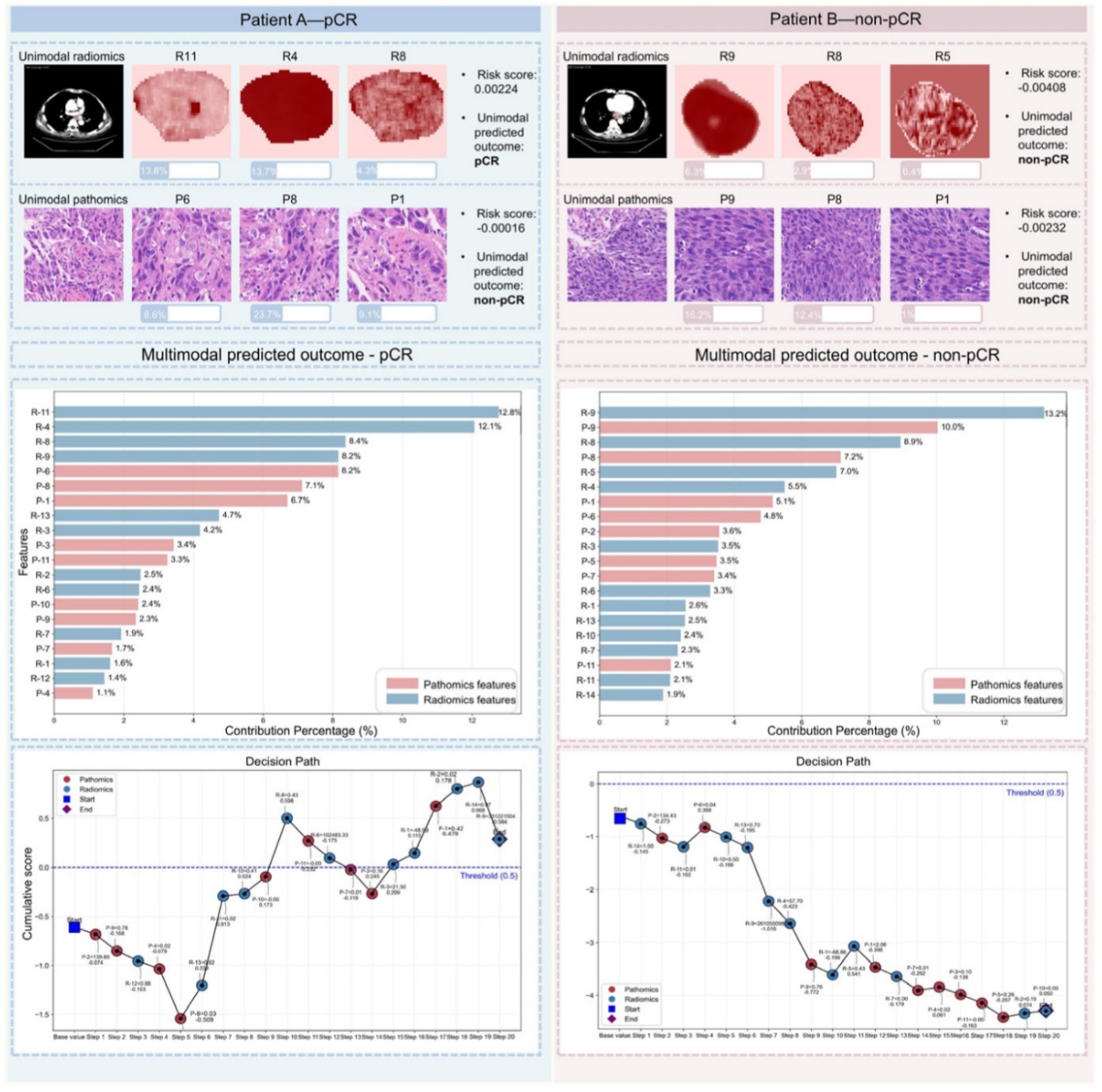
“Interpretable multimodal radiopathomics model predicting pathological complete response to neoadjuvant chemoimmunotherapy in esophageal squamous cell carcinoma”
Published in Journal for ImmunoTherapy of Cancer on 5 December 2025 and selected as a monthly spotlight, the study led by Prof. Yongling JI and Prof. Zhen ZHANG developed an interpretable multimodal machine learning model integrating pretreatment CT radiomics and H&E-stained whole-slide pathology features. The intermediate fusion model achieved AUCs of 0.97, 0.78, and 0.76 in training, internal, and external test sets, outperforming single-modality models. Model interpretability was prioritized through SHAP analysis, case-level contribution maps, and decision path visualization, enabling clinicians to trace how the model weighs evidence when predictions differ. A user-friendly graphical interface supports potential clinical translation for predicting pathological complete response in esophageal squamous cell carcinoma patients undergoing neoadjuvant chemoimmunotherapy.